- Tools
- Downloads
- Links
- Community
- Species
- About
- Help
hernandez
two closely linked genes coding for pipsqueak domain proteins - effector genes that act downstream of homothorax and Distal-less to control differentiation of distal antennal structures
Please see the JBrowse view of Dmel\danr for information on other features
To submit a correction to a gene model please use the Contact FlyBase form
AlphaFold produces a per-residue confidence score (pLDDT) between 0 and 100. Some regions with low pLDDT may be unstructured in isolation.
Gene model reviewed during 5.49
Gene model reviewed during 5.55
There is only one protein coding transcript and one polypeptide associated with this gene
Interacts with itself, dan, ey and dac to form a complex (or complexes) containing the RD factors.
Click to get a list of regulatory features (enhancers, TFBS, etc.) and gene disruptions (point mutations, indels, etc.) within or overlapping Dmel\danr using the Feature Mapper tool.
The testis specificity index was calculated from modENCODE tissue expression data by Vedelek et al., 2018 to indicate the degree of testis enrichment compared to other tissues. Scores range from -2.52 (underrepresented) to 5.2 (very high testis bias).
Comment: anlage in statu nascendi
Comment: anlage in statu nascendi
Comment: anlage in statu nascendi
Comment: anlage in statu nascendi
Comment: anlage in statu nascendi
Comment: reported as procephalic ectoderm anlage in statu nascendi
Comment: reported as procephalic ectoderm anlage in statu nascendi
Comment: reported as procephalic ectoderm anlage in statu nascendi
Comment: reported as procephalic ectoderm anlage
Comment: reported as procephalic ectoderm anlage
Comment: reported as procephalic ectoderm anlage
Comment: reported as procephalic ectoderm anlage
Comment: reported as procephalic ectoderm primordium
Comment: reported as procephalic ectoderm primordium
Comment: reported as procephalic ectoderm primordium
Comment: reported as procephalic ectoderm primordium
Comment: reported as procephalic ectoderm primordium
Comment: reported as procephalic ectoderm primordium
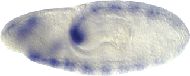
danr transcript is undetectable in embryos through the syncytial blastoderm stage, suggesting that it is not maternally supplied. In stage 5 embryos danr transcript is expressed uniformly, except in the terminal regions, and in the ventral part of the embryo corresponding to the mesoderm anlage. There is also a dorsal cap of expression at the anterior of the stage 5 embryo. Expression becomes ventralized at stage 8, concentrating in the ventral neurectoderm and decreasing in the lateral ectoderm. Expression is confined to the ventral nerve cord from embryonic stage 11-15.
Expression of the dan and danr transcripts in imaginal discs is restricted to the eye and antennal discs. Expression is strong anterior to the morphogenetic furrow with weaker expression posterior to the furrow. Strong signal is also observed in a ring corresponding to the region of the third antennal segment and in the central region of the antennal primordia. This expression pattern persists in early pupae.
JBrowse - Visual display of RNA-Seq signals
View Dmel\danr in JBrowse



3-87
3-85.6
Please Note FlyBase no longer curates genomic clone accessions so this list may not be complete
Please Note This section lists cDNAs and ESTs that fall within the genomic extent of the gene model, which may include cDNAs and ESTs of genes within introns, or of overlapping genes. Please see JBrowse for alignment of the cDNAs and ESTs to the gene model.
For each fully sequenced cDNA the DGRC maintains various forms of the cDNA (e.g tagged or untagged) in several different host vectors for subsequent cloning and expression in Drosophila and Drosophila cell lines.
Source for identity of: danr CG13651